SPL Smart Protein Layers
Kit for normalization of Western blots using total protein
The Smart Protein Layers (SPL) kit enables reliable target normalization and quantification. Fluorescent SPL Label binds to all proteins in the sample prior to electrophoresis and serves as a reliable total protein indicator to normalize your target protein. Corresponding bi-fluorescent SPL Standards monitor labeling efficiency, provide additional information about the sample and enable automated data evaluation.
With SPL the determined expression of a specific target protein present on the blot can be normalized to its corresponding sample to provide reliable and precise comparison of fold changes between samples.
SPL Features
- • highly sensitive protein detection in gels and blots - stainfree
- • normalization of the protein content bewteen samples
- • step-by-step monitoring of the Western Blot workflow
- • precise quantification of target protein expression
- • accurate comparison of data between different experiments
.
.
Depending on your way of target detection and fluorescence imaging capabilities, we offer different SPL kits.
SPL Kits
| Product No. | Description | Size | Recommended
Target Detection |
Required
Fluorescence Imaging |
Price |
| PR913 |
SPL Red Kit 20W
|
200 rcts. | Immuno Blue
or ECL |
blue + red | 189,00 EUR |
| PR926 |
SPL Red Kit 40W
|
400 rcts. | Immuno Blue
or ECL |
blue + red | 329,00 EUR |
| PR916 |
SPL Blue Kit 20W
|
200 rcts. | Red-fluoresc. AB
|
blue + red | 189,00 EUR |
| PR925 |
SPL Blue Kit 40W
|
400 rcts. | Red-fluoresc. AB
|
blue + red | 329,00 EUR |
| PR917 |
SPL Red-IR Kit 20W
|
200 rcts. | IR-fluoresc. AB
or ECL |
red + IR | 189,00 EUR |
| PR927 |
SPL Red-IR Kit 40W
|
400 rcts. | IR-fluoresc. AB
or ECL |
red + IR | 329,00 EUR |
.
SPL Technology
Using different distinct fluorescent wavelenghts, each providing information. That is the background of the patented SPL Smart Protein Layers Technology. The most important additonal information for Western Blot analysis is: How much sample protein is present on the blot when a target protein out of this sample is detected? A comparison between expressed target proteins of different samples is valid only (quantitative Western Blot analysis), when the amount of present total sample protein is equal or can be normalized to the target signal.
However, the SPL technology provides more information than that. SPL measures potential differences in total protein content and it can tell you the reason for the differences. SPL also monitors the daily performance of the secondary antibody you are using.
The backbone of the SPL Smart Protein Layers technology is provided by a combination of fluorescent sample standards (Smartalyzers) and a rapid and controlled sample + Smartalyzer labeling by specific fluorophores (Smart Labels).
.
WB Normalization using "House-keeping proteins"
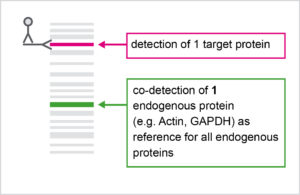
.
WB Normalization using SPL
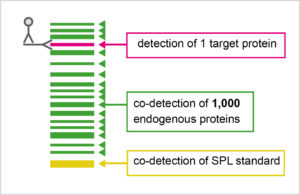
The SPL Kit is composed out of three components:
- • The Smartalyzer is a part of the SPL Loading Buffer added to the sample prior heating and SDS-PAGE. This sample dependent fluorescent standard allows for the precise normalization and quantifiaction.
- • Fluorescent Smart Label added to the sample prior heating (5min 37-95 degress centigrade) and SDS-PAGE allow for the highly sensitive visualization (1 ng or less) of proteins during the entire Western Blot analysis.
- • The Calibrator is a gel and blot specific standard for the comparison of data derived from different experiments. Furthermore, theCalibrator is also a fluorescent proteinladder.
Benefit
- • Normalization of 1 Western Blot target protein to approx. 800 - 1,000 corresponding endogenous proteins
Additional Benefit
- • Stain-free protein visualization
(sensitivity of silver but 1,000 higher dynamic range) - • Protein load equalizer
(normalization of samples with unequal protein content) - • Precise protein expression quantification
(precise lane-to-lane, gel-to-gel and blot-to-blot comparisons)
.
Related Documents
Publications
Rausche J, Stenzel I, Stauder R et al. A phosphoinositide 5‐phosphatase from Solanum tuberosum is activated by PAMP‐treatment and may antagonize phosphatidylinositol 4,5‐bisphosphate at Phytophthora infestans infection sites. New Phytol. 2020, doi:10.1111/nph.16853
More publications and selected customers you will find here.
Manuals
PDF Quick Guide SPL Labeling (Hands-on guide for the lab bench)
.
PDF Manual PR916/PR925 SPL Blue Kit 20W / 40W
.
PDF Manual PR913/PR926 SPL Red Kit 20W / 40W
.
PDF Manual PR917/PR927 SPL Red-IR Kit 20W / 40W
Safety data sheet
PDF Safety data sheet PR917/PR927 SPL Kits for Western Blots
.
Evaluation Software
LabImage 1D SPL
Standard for Blot-to-Blot Comparison
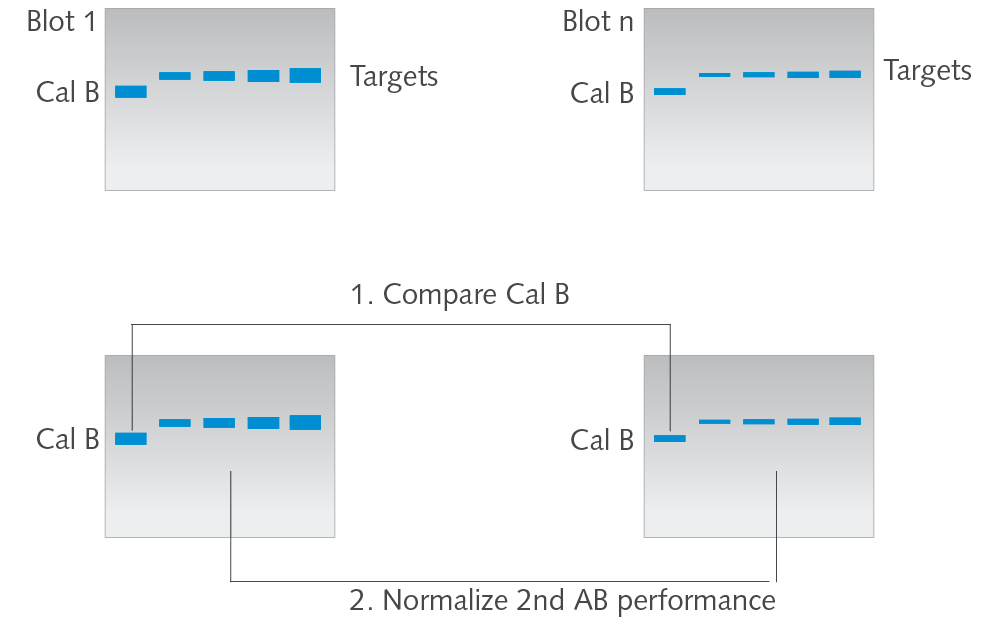
Calibrator B
The Calibrator B (50 kDa, either mouse, rabbit or goat) is specific to secondary antibody (either anti-mouse, anti-rabbit or anti-goat). Cal B therefore allows to check for the performance of the secondary AB. Furthermore, when applied to the blot, Cal B allows for precise blot-to-blot analysis by normalization of secondary AB signal intensities.
The kit contains 1 vial Cal B sufficient for 20 Western Blot experiments (1D) or 10 2D-Western Blots.
.
| Product No. | Description | Price |
| PR921 | Calibrator B
Mouse |
50,00 EUR |
| PR922 | Calibrator B
Rabbit |
50,00 EUR |
| PR923 | Calibrator B
Goat |
50,00 EUR |
.
.
.
Frequently asked questions (FAQs)
.
Does the SPL labels interfere with antibody recognition?
The SPL technology uses the labeling chemistry of the so-called minimal lysine labeling. Only 2-3% of each protein is getting labeled with one fluorophore per protein. A lysine-residue important for antibody recognition can be conjugated with fluorophore leading to antibody interference. However, since 97-98% of this protein is not labeled, it does not matter.
Do differences in sample labeling efficiency lead to false protein detection signals?
The minimal labeling chemistry was originally developed for purified protein samples for 2D gel analysis (Refraction-2D, 2D-DIGE). For 1D gel analysis and Western Blots the protein preparation is more crude: proteins are extracted from proteins and, after the addition of loading buffer separated by SDS-PAGE.However, not purified protein samples may contain components leading to a decrease in labeling efficiency as this is a chemical reaction. Special compounds in the SPL loading buffer ensure optimal labeling conditions for most samples. In addition, the efficiency of the SPL labeling of each sample is carefully monitored and if necessary normalized by the control standard (the Smartalyzer, patented technology).
What is the difference between a (fluorescent) stain and a Smart Label?
A stain is a dye binding non-covalently to proteins. The staining procedure includes the staining and than several destaining steps. The reproducibility of stains is related to the staining efficiency (e.g. time, temperature) and may suffer from background.
Smart Labels are fluorescent dyes, which bind covalently to proteins. Smart labeled proteins can be visualized directly after electrophoresis – no staining or destaining steps are required. Smart Labels are highly sensitive with a wide dynamic range.
.
Is the Smart Labeling exclusive for certain proteins or will it favor certain proteins?
Smart Labels bind to all proteins in a non-saturated way to allow for precise quantification. The labeling chemistry and ratio of labels to proteins are well established (see more than 1,000 DIGE and Refraction-2D publications) and guarantees an even labeling of all proteins within one sample.
.
What may inhibit the labeling process?
Due to special SPL buffers, the Smart Label reaction is very robust. Smart Labeling is compatible with all commonly used buffer systems. However, high concentrations of reducing agents (> 15 mM DTT or > 10 % beta-mercaptoethanol) and amines > 400 mM may interfere with labeling reaction. The SPL Smartalyzer (SMA) is added to the labeling reaction as a control. Its unique design monitors labeling efficiency and if necessary compensates for unequal labeling.
.
For sample Smart Labeling, what is the maximum and minimum volume of my sample? The maximum volume is 10 µl per Smart Label reaction (containing up to 100 µg of protein). The minimum volume is only given by what you can still handle.
.
If I have to dilute my protein sample before Smart Labeling, can I use my own buffer? Yes please.
.
Why is the Smartalyzer (SMA) bi-fluorescent?
The SMA itself is mono-fluorescent (e.g. SMA basic blue). SMA is added to the protein sample before Smart Labeling. SMA is labeled together with the protein sample (e.g. with Smart Label Red). So SMA becomes bi-fluorescent (e.g. basic blue and label red). SMA basic allows for normalization of gel loading and SMA Label allows for normalization of labeling.
.
Which size of the standard SMA should I use, 12.5 kDa or 80 kDa?
The size of the SMA should differ from the size of your target protein and highly abundant proteins within your sample.
.
How do I avoid that the SMA size S (12.5 kDa) runs out of the gel?
Stop electrophoresis before the dye front runs out of the gel. Cut the dye front manually before fluorescence imaging (dye front may interfere with Imaging).
.
Does the Smart Label interfere with subsequent Mass Spectrometry?
No. Smart Labels do not effect enzymatic digestions nor sequence coverage.
.
Does the Smart Label interfere with antibody recognition?
No. Only one amino acid of a minimal amount of protein is labeled.
.
Can I stain Smart labeled gels and then cut out bands?
Yes. Smart Labels are compatible with all commonly visual protein stains (e.g. Coomassie® or silver nitrate) and subsequent protein identification by mass spectrometry.
.
SPL User






